Plant Systematics
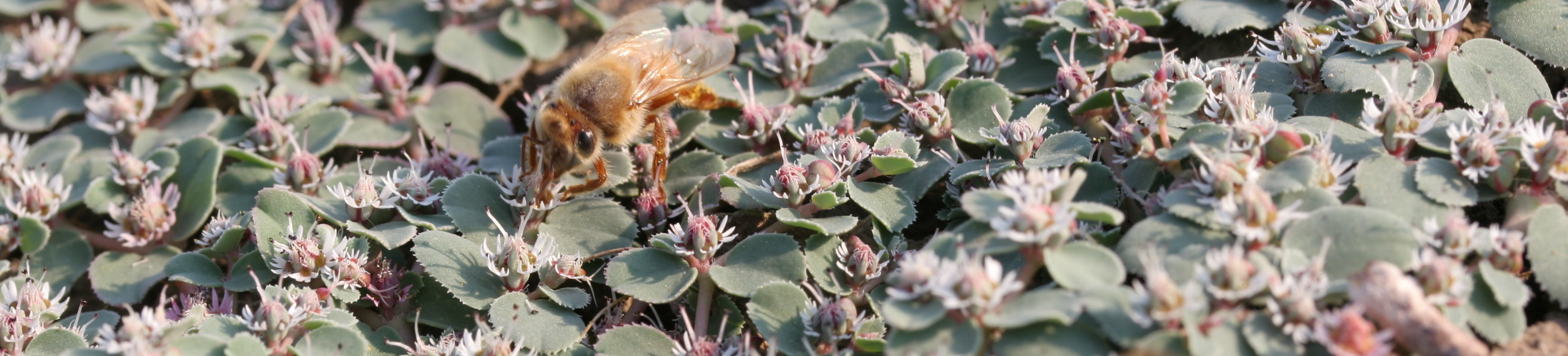
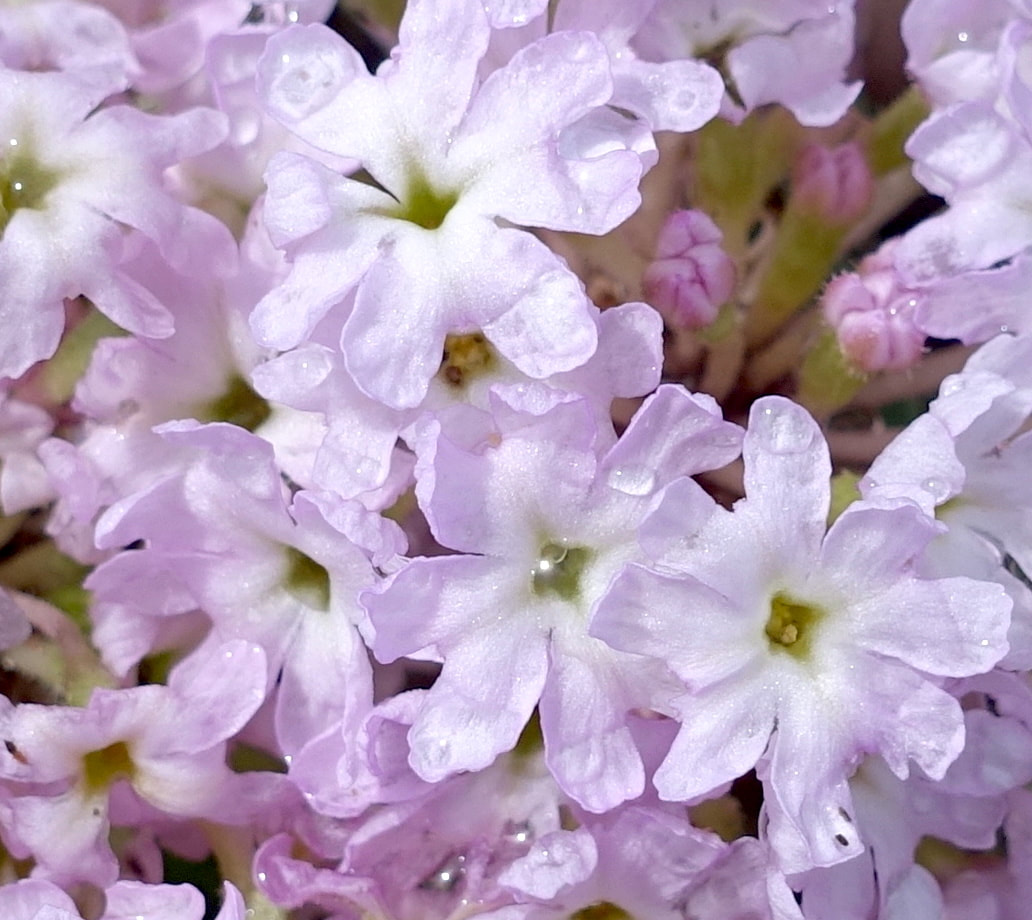
We are interested in systematics, biogeography, and lineage diversification in various plant groups, such as Euphorbia and the Caryophyllales.
Morales-Briones, D.F., N. Lin, E.Y. Huang, D.L. Grossenbacher, J.M. Sobel, C.D. Gilmore, D.C. Tank, Y. Yang. 2022. Phylogenomic analyses in Phrymaceae reveal extensive gene tree discordance in relationships among major clades. American Journal of Botany. 109(6): 1035–1064 Open Access
Morales-Briones, D.F., B. Gehrke, C.-H. Huang, A. Liston, H. Ma, H.E. Marx, D.C. Tank, Y. Yang. 2022. Analysis of paralogs in target enrichment data pinpoints multiple ancient polyploidy events in Alchemilla s.l. (Rosaceae). Systematic Biology. 71(1):190–207. Open access
Morales-Briones, D.F., G. Kadereit, D.T. Tefarikis, M.J. Moore, S.A. Smith, S.F. Brockington, A. Timoneda, W.C. Yim, J.C. Cushman, and Y. Yang. Disentangling sources of gene tree discordance in phylogenomic datasets: Testing ancient hybridizations in Amaranthaceae s.l. Systematic Biology. 70(2):219–235. Open access
Yang, Y., C.W. Morden, M.J. Sporck-Koehler, L. Sack, W.L. Wagner, and P.E. Berry. 2018. Repeated range expansion and niche shift in a volcanic hotspot archipelago: radiation of C4 Hawaiian Euphorbia (Euphorbiaceae). Ecology and Evolution. doi:10.1002/ece3.4354
Yang, Y., R. Riina, J.J. Morawetz, T. Haevermans, X. Aubriot and P.E. Berry. 2012. Molecular phylogenetics and classification of Euphorbia subgenus Chamaesyce: a group with high diversity in photosynthetic systems and growth forms. Taxon 61(4):764–789
Yang, Y. and P.E. Berry. 2011. Phylogenetics of the Chamaesyce clade (Euphorbia, Euphorbiaceae): reticulate evolution and long-distance dispersal in a prominent C4 lineage. American Journal of Botany 98(9):1486–1503
Morales-Briones, D.F., B. Gehrke, C.-H. Huang, A. Liston, H. Ma, H.E. Marx, D.C. Tank, Y. Yang. 2022. Analysis of paralogs in target enrichment data pinpoints multiple ancient polyploidy events in Alchemilla s.l. (Rosaceae). Systematic Biology. 71(1):190–207. Open access
Morales-Briones, D.F., G. Kadereit, D.T. Tefarikis, M.J. Moore, S.A. Smith, S.F. Brockington, A. Timoneda, W.C. Yim, J.C. Cushman, and Y. Yang. Disentangling sources of gene tree discordance in phylogenomic datasets: Testing ancient hybridizations in Amaranthaceae s.l. Systematic Biology. 70(2):219–235. Open access
Yang, Y., C.W. Morden, M.J. Sporck-Koehler, L. Sack, W.L. Wagner, and P.E. Berry. 2018. Repeated range expansion and niche shift in a volcanic hotspot archipelago: radiation of C4 Hawaiian Euphorbia (Euphorbiaceae). Ecology and Evolution. doi:10.1002/ece3.4354
Yang, Y., R. Riina, J.J. Morawetz, T. Haevermans, X. Aubriot and P.E. Berry. 2012. Molecular phylogenetics and classification of Euphorbia subgenus Chamaesyce: a group with high diversity in photosynthetic systems and growth forms. Taxon 61(4):764–789
Yang, Y. and P.E. Berry. 2011. Phylogenetics of the Chamaesyce clade (Euphorbia, Euphorbiaceae): reticulate evolution and long-distance dispersal in a prominent C4 lineage. American Journal of Botany 98(9):1486–1503
Functional Phylogenomics
We develope methods for mining genomic and transcriptomic data at the macroevolutionary scale that go beyond "single-copy genes". By doing so now we are able to detect the phylogenetic and functional locations of gene tree discordance, gene and genome duplication, molecular rate heterogeneity, and natural selection at a scale that is never before possible.
Representative publications:
McKain, M.R., M.G. Johnson, S. Uribe‐Convers, D. Eaton, and Y. Yang. 2018. Practical considerations for plant phylogenomics. Applications in Plant Sciences 6(3): e1038.
Yang, Y., M.J. Moore, S.F. Brockington, J. Mikenas, J. Olivieri, J.F. Walker, and S.A. Smith. 2018. Improved transcriptome sampling pinpoints 26 ancient and more recent polyploidy events in Caryophyllales, including two allopolyploidy events. New Phytologist. 217(2): 855–870
Yang, Y., M.J. Moore, S.F. Brockington, D.E. Soltis, G.K.-S. Wong, E.J. Carpenter, Y. Zhang, L. Chen, Z.-X. Yan, Y.-L. Xie, R.F. Sage, S. Covshoff, J.M. Hibberd, M.N. Nelson, and S.A. Smith. 2015. Dissecting molecular evolution in the highly diverse plant clade Caryophyllales using transcriptome sequencing. Molecular Biology and Evolution. 32(8): 2001-2014.
Yang, Y. and S.A. Smith. 2014. Orthology inference in non-model organisms using transcriptomes and low-coverage genomes: improving accuracy and matrix occupancy for phylogenomics. Molecular Biology and Evolution. 31 (11): 3081–3092.
McKain, M.R., M.G. Johnson, S. Uribe‐Convers, D. Eaton, and Y. Yang. 2018. Practical considerations for plant phylogenomics. Applications in Plant Sciences 6(3): e1038.
Yang, Y., M.J. Moore, S.F. Brockington, J. Mikenas, J. Olivieri, J.F. Walker, and S.A. Smith. 2018. Improved transcriptome sampling pinpoints 26 ancient and more recent polyploidy events in Caryophyllales, including two allopolyploidy events. New Phytologist. 217(2): 855–870
Yang, Y., M.J. Moore, S.F. Brockington, D.E. Soltis, G.K.-S. Wong, E.J. Carpenter, Y. Zhang, L. Chen, Z.-X. Yan, Y.-L. Xie, R.F. Sage, S. Covshoff, J.M. Hibberd, M.N. Nelson, and S.A. Smith. 2015. Dissecting molecular evolution in the highly diverse plant clade Caryophyllales using transcriptome sequencing. Molecular Biology and Evolution. 32(8): 2001-2014.
Yang, Y. and S.A. Smith. 2014. Orthology inference in non-model organisms using transcriptomes and low-coverage genomes: improving accuracy and matrix occupancy for phylogenomics. Molecular Biology and Evolution. 31 (11): 3081–3092.
Evolution of gene families and modules
|
What are the mechanisms underly trait evolution?
Gene families and networks that we are investigating include those involved in centromeres, stress tolerance, and evolution of metabolites. The work is currently funded by the National Science Foundation (NSF) through two awards: NSF award on Caryophyllales metobolite evolution NSF-funded project on spectral biology Representative publications: Tefarikis, D.T., D.F. Morales-Briones, Y. Yang, G. Edwards, G. Kadereit. 2022. On the hybrid origin of the C2 Salsola divaricata agg. (Amaranthaceae) from C3 and C4 parental lineages. New Phytologist. 234:1876–1890. Open access Lopez-Nievesa, S., Y. Yang, A. Timoneda, M.-M. Wang, T. Feng, S.A. Smith, S.F. Brockington, and H.A. Maeda. 2018. Relaxation of tyrosine pathway regulation underlies the evolution of betalain pigmentation in Caryophyllales. New Phytologist 217(2): 896–908 Brockington S.F., Y. Yang, F. Gandia-Herrero, S. Covshoff, J.M. Hibberd, R.F. Sage, G.K.-S. Wong, M.J. Moore and S.A. Smith. 2015. Lineage specific gene radiations underly the evolution of novel betalain pigmentation in Caryophyllales. New Phytologist. 207(4) 1170–1180 |